sRNAbench Results for job ID: ZAPFTBMPBGCPIDM
The results will be stored for 15 days (This will be removed on: Tue 14 May 2019).
If you use the sRNAbench webserver plese check How to Cite.
Download all results as zip file here
Input File:SRR1563060
---- Mapping parameters and Annotations ----
Number of allowed mismatches: 1 (mm=1)
Alignment type: seed alignment (alignType=n)
Length of the seed: 20 (seed=20)
Used genome assembly bowtie1 index: GRCh38_p10_mp:NC_007605
---- ANNOTATIONS ----
Used miRNAs reference species from miRBase v22
Used miRNAs for species: hsa:ebv
Used ncRNA annotations:
libs=GRCh38_p10_genomic_tRNA (Genomic tRNA database)
libs=hsa_yRNA (yRNAs)
libs=vRNA (vault RNAs)
libs=GRCh38_p10_ncRNA (Ensembl release 91 (ncRNA))
libs=GRCh38_p10_cdna (Ensembl release 91 (cDNA))
libs=GRCh38_p10_RNAcentral (RNAcentral database 7.0)
---- Preprocessing and adapter trimming parameters: ----
Unknown protocol: Illumina
---No quality filtering was used (only reads with ambigous bases are filtered out- ---
Number of Raw input reads : 6203008
Number of adapter trimmed reads: 6114811 (98.57815756484595)
--- Number of filtered reads ----
Quality Filtered Reads : 18872
Reads below min read count : 309386
Reads above max. length : 0
Reads below min. length : 1457621
Percentage of AdapterDimer : 3.4880611813859517
Reads in analysis: 4328932 (69.7876256164751)
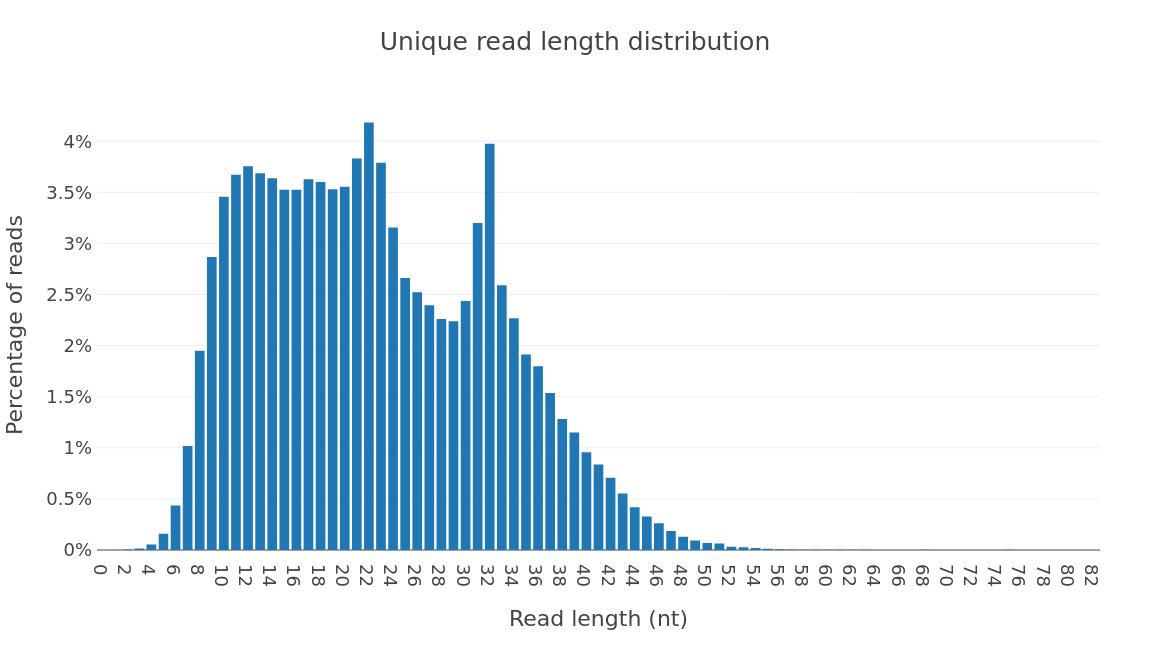
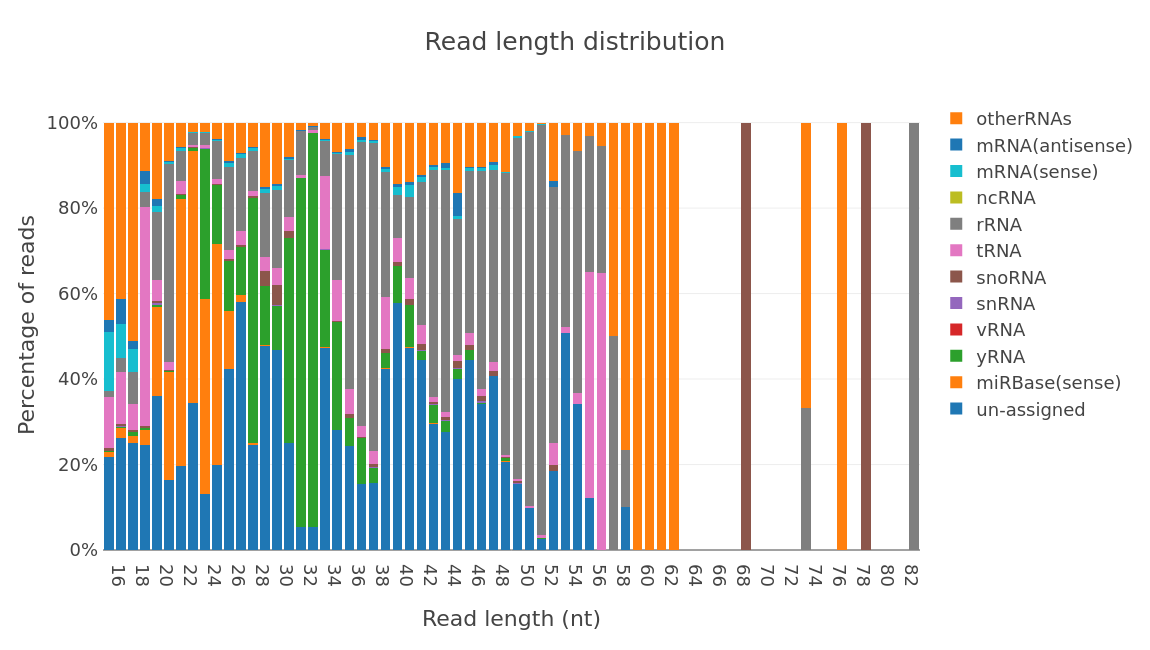
Profiling results by rna type.
| Library | Description | Orientation | UR | URperc | RC | RCperc | RPMassigned | RPManalysis |
|---|---|---|---|---|---|---|---|---|
| mature | miRBase v22 | antisense | 2 | 0 | 4 | 0 | 1.17 | 1.05 |
| hsa_yRNA | yRNAs | sense | 3678 | 4.62 | 1461807 | 38.44 | 427985.2 | 384408.06 |
| GRCh38_p10_genomic_tRNA | Genomic tRNA database | sense | 7175 | 9.01 | 211344 | 5.56 | 61876.91 | 55576.65 |
| GRCh38_p10_RNAcentral | RNAcentral database 7.0 | sense | 3499 | 4.39 | 498036 | 13.1 | 145814.08 | 130967.4 |
| GRCh38_p10_ncRNA | Ensembl release 91 (ncRNA) | antisense | 7549 | 9.48 | 108827 | 2.86 | 31862.17 | 28617.99 |
| GRCh38_p10_ncRNA | Ensembl release 91 (ncRNA) | sense | 11299 | 14.19 | 150548 | 3.96 | 44077.17 | 39589.27 |
| GRCh38_p10_genomic_tRNA | Genomic tRNA database | antisense | 4 | 0.01 | 13 | 0 | 3.81 | 3.42 |
| hairpin | -- | sense | 180 | 0.23 | 2383 | 0.06 | 697.69 | 626.65 |
| GRCh38_p10_cdna | Ensembl release 91 (cDNA) | sense | 5446 | 6.84 | 111771 | 2.94 | 32724.11 | 29392.17 |
| GRCh38_p10_cdna | Ensembl release 91 (cDNA) | antisense | 3934 | 4.94 | 62364 | 1.64 | 18258.82 | 16399.72 |
| hairpin | -- | antisense | 7 | 0.01 | 16 | 0 | 4.68 | 4.21 |
| mature | miRBase v22 | sense | 6434 | 8.08 | 788123 | 20.73 | 230745.22 | 207250.91 |
| vRNA | vault RNAs | sense | 308 | 0.39 | 7453 | 0.2 | 2182.08 | 1959.9 |
| GRCh38_p10_RNAcentral | RNAcentral database 7.0 | antisense | 247 | 0.31 | 12866 | 0.34 | 3766.88 | 3383.34 |
| un-assigned | -- | --- | 29878 | 37.52 | 387193 | 10.18 | 0 | 101819.26 |
Genome/Library mapping results
- Unique Genome mapped reads:
- 79640(88.25%)
- Genome mapped reads:
- 3802748(87.84%)
Download BedGraph data for UCSC/Ensemble browser
| Description | Link |
|---|---|
| Description: Regions with mapped reads in bed format (the score shows gives the maximum expression value in the region). ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All genome mapped reads in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All reads mapped to the (+) strand in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All reads mapped to the (-) strand in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: Regions with mapped reads >=19nt and <=23nt in bed format (the score shows gives the maximum expression value in the region). ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: Genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: (+) strand genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: (-) strand genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: Regions with mapped reads in bed format (the score shows gives the maximum expression value in the region). ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All genome mapped reads in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All reads mapped to the (+) strand in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: All reads mapped to the (-) strand in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: Regions with mapped reads >=19nt and <=23nt in bed format (the score shows gives the maximum expression value in the region). ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: Genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: (+) strand genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
| Description: (-) strand genome mapped reads >=19nt and <=23nt in BedGraph format ;Multiple mapping treatment: Full assignment (full read count in all genome possitions) | Download data |
MiR profiling results
- Reads mapped to miRNAs:
- 788123 (23.07%)
- Reads mapped to miRBase hairpins:
- 2383 (0.07%)
- Detected hairpin miR:
- 393 (20.24%)
- Detected mature miR:
- 566 (20.96%)
- MicroRNA details:
- isomiR details:
Ensembl release 91 (ncRNA)
- Mapped reads in sense direction:
- 150548(4.41%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 108827(3.19%)
- MA antisense
- SA antisense
vault RNAs
- Mapped reads in sense direction:
- 7453(0.22%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 0(0.0%)
- MA antisense
- SA antisense
RNAcentral database 7.0
- Mapped reads in sense direction:
- 498036(14.58%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 12866(0.38%)
- MA antisense
- SA antisense
Ensembl release 91 (cDNA)
- Mapped reads in sense direction:
- 111771(3.27%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 62364(1.83%)
- MA antisense
- SA antisense
yRNAs
- Mapped reads in sense direction:
- 1461807(42.8%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 0(0.0%)
- MA antisense
- SA antisense
Genomic tRNA database
- Mapped reads in sense direction:
- 211344(6.19%)
- multiple assignment (MA)
- single assignment (SA)
- Mapped reads in antisense direction:
- 13(0.0%)
- MA antisense
- SA antisense
- tRNA genes:
- details